pacman::p_load(spdep, sp, tmap, sf, ClustGeo, cluster, factoextra, NbClust, tidyverse, GGally)In Class exercise 10
6.0 Loading the R packages
shan_sf <- read_rds("data/In-class_Ex09/rds/shan_sf.rds")
shan_ict <- read_rds("data/In-class_Ex09/rds/shan_ict.rds")
shan_sf_cluster <- read_rds("data/In-class_Ex09/rds/shan_sf_cluster.rds")6.1 Conventional Hierarchical Clustering
In R, many packages provide functions to calculate distance matrix. We will compute the proximity matrix by using dist() of R.
dist() supports six distance proximity calculations, they are: euclidean, maximum, manhattan, canberra, binary and minkowski. The default is euclidean proximity matrix.
The code chunk below is used to compute the proximity matrix using euclidean method.
proxmat <- dist(shan_ict, method = "euclidean")
hclust_ward <- hclust(proxmat, method = "ward.D")
groups <- as.factor(cutree(hclust_ward, k=6))hclust() will take the proximity matrix to perform hierarchical clustering to create a hierarchical clustering object to get the the groups based on the cutree( method
This chunk of code is meant to tidy the shan_sf_cluster dataset
shan_sf_cluster <- cbind(shan_sf, as.matrix(groups)) %>%
rename(`CLUSTER`=`as.matrix.groups.`) %>%
select(-c(3:4, 7:9)) %>%
rename(TS = TS.x)This chunk of code to create the dendogram
plot(hclust_ward, cex = 0.6)
rect.hclust(hclust_ward, k = 6, border = 2.5)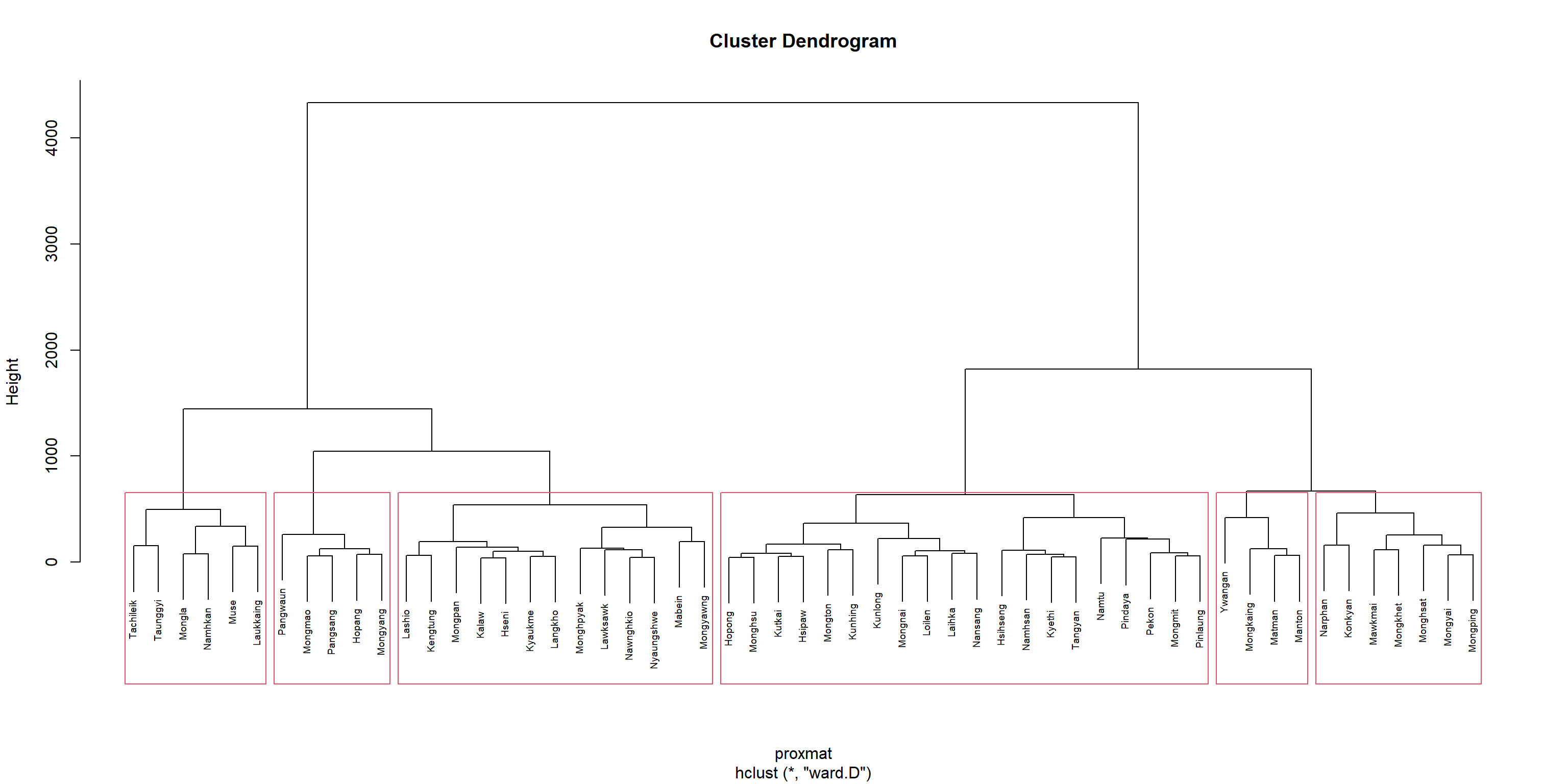
This chunk of code to create the cluster map of the shan_sf_cluster object
qtm(shan_sf_cluster, "CLUSTER")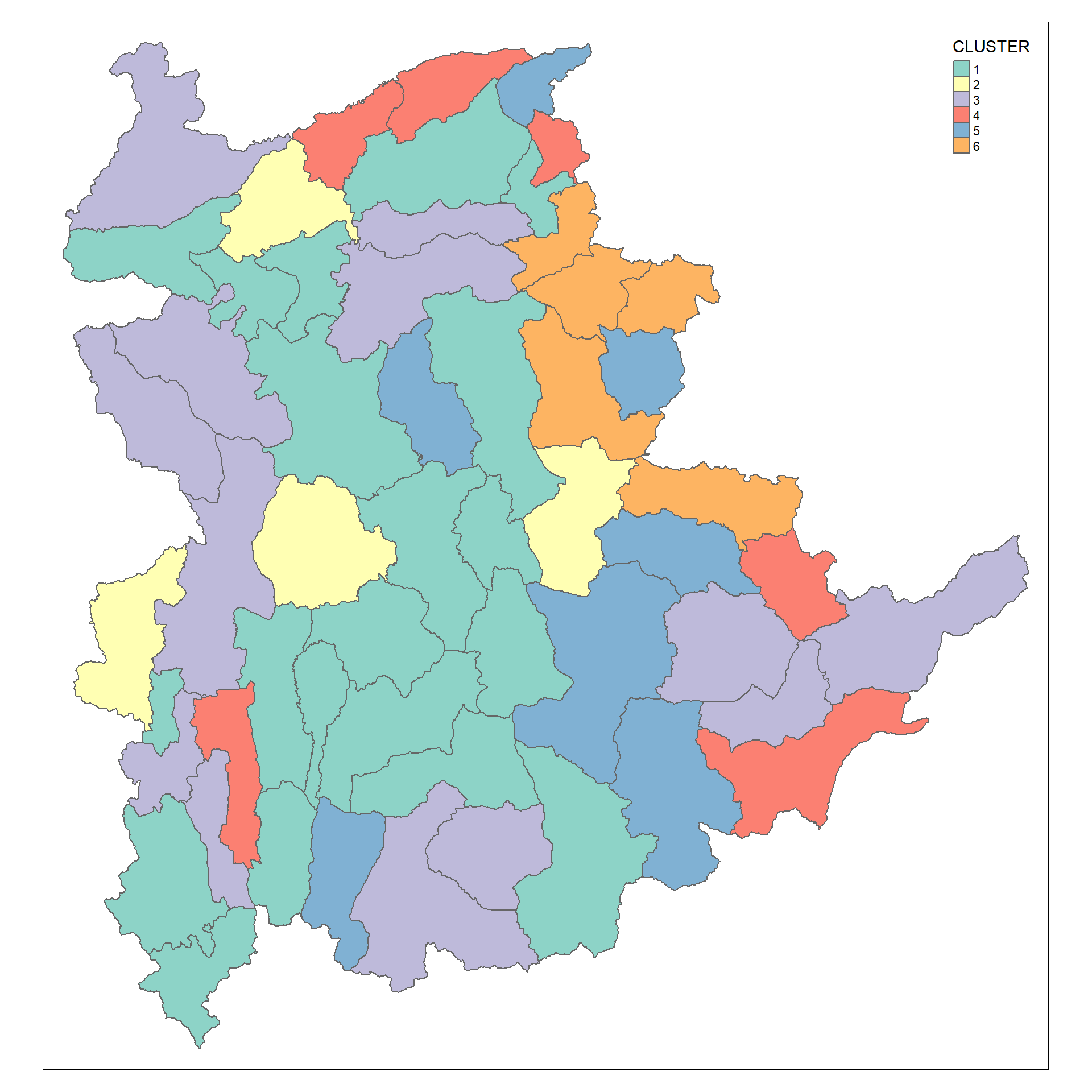
Spatially Constrained Clustering
SKATER (Spatial ’K’luser Analysis by Tree Edge Removal) Alogrithm
REDCAP (Reorganisation with dynamically
ClustGeo Algorithm
SKATER Algorithm
Spatially Constrained Clustering: SKATER Method
- Computing nearest neighbours (Minimum Spanning Tree)
shan.nb <- poly2nb(shan_sf)
summary(shan.nb)Neighbour list object:
Number of regions: 55
Number of nonzero links: 264
Percentage nonzero weights: 8.727273
Average number of links: 4.8
Link number distribution:
2 3 4 5 6 7 8 9
5 9 7 21 4 3 5 1
5 least connected regions:
3 5 7 9 47 with 2 links
1 most connected region:
8 with 9 links- Visualising the neighbours
plot(st_geometry(shan_sf),
border=grey(.5))
pts <- st_coordinates(st_centroid(shan_sf))
plot(shan.nb, pts, col="blue", add=TRUE)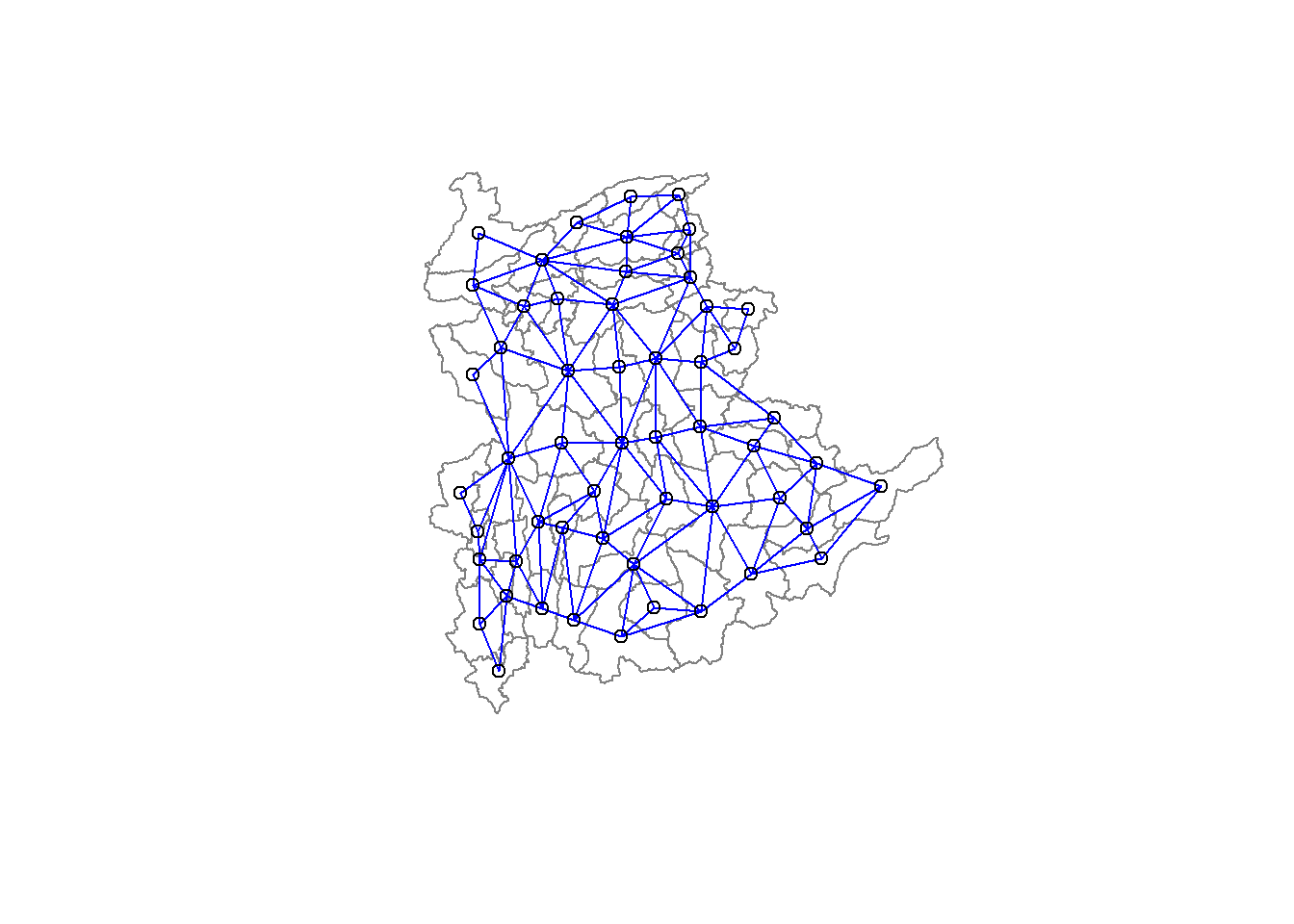
- Computing minimum spanning tree (MST)
- Calculating edge costs
lcosts <- nbcosts(shan.nb, shan_ict)- Incorporating these costs into a weights object
shan.w <- nb2listw(shan.nb, lcosts, style = "B")
summary(shan.w)Characteristics of weights list object:
Neighbour list object:
Number of regions: 55
Number of nonzero links: 264
Percentage nonzero weights: 8.727273
Average number of links: 4.8
Link number distribution:
2 3 4 5 6 7 8 9
5 9 7 21 4 3 5 1
5 least connected regions:
3 5 7 9 47 with 2 links
1 most connected region:
8 with 9 links
Weights style: B
Weights constants summary:
n nn S0 S1 S2
B 55 3025 76267.65 58260785 522016004- Visualising MST
shan.mst <- mstree(shan.w)plot(st_geometry(shan_sf), border=gray(.5))
plot.mst(shan.mst,
pts,
col="blue",
cex.lab=0.7,
cex.circles = 0.005,
add=TRUE)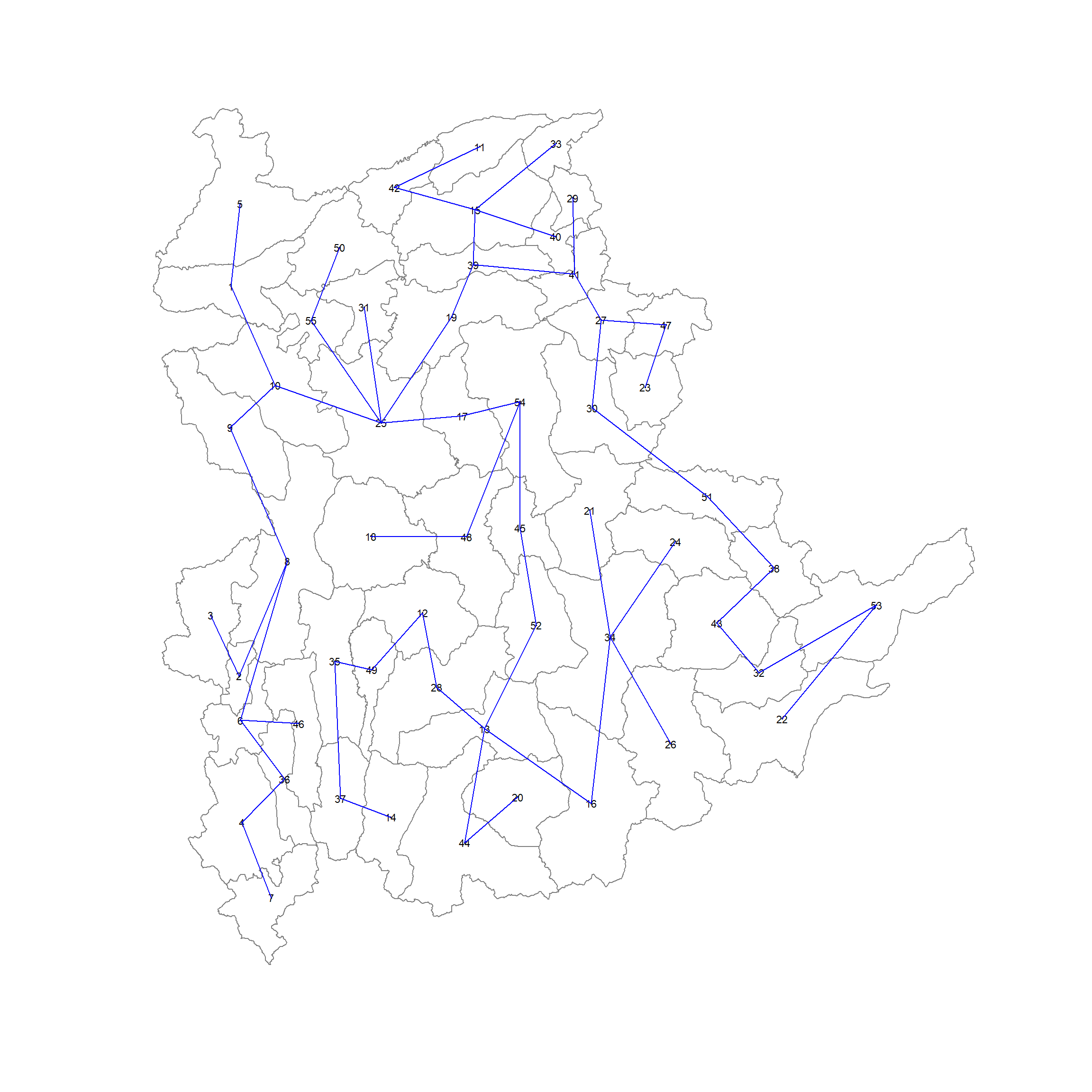
Computing spatially constrained clusters using SKATER method
skater.clust6 <- skater(edges = shan.mst[,1:2],
data = shan_ict,
method = "euclidean",
ncuts = 5)The following code chunk plots the skater tree
plot(st_geometry(shan_sf), border=gray(.5))
plot(skater.clust6,
pts,
cex.lab=.7,
groups.colors=c("red", "green", "blue", "brown", "pink"),
cex.circles = 0.005,
add=TRUE)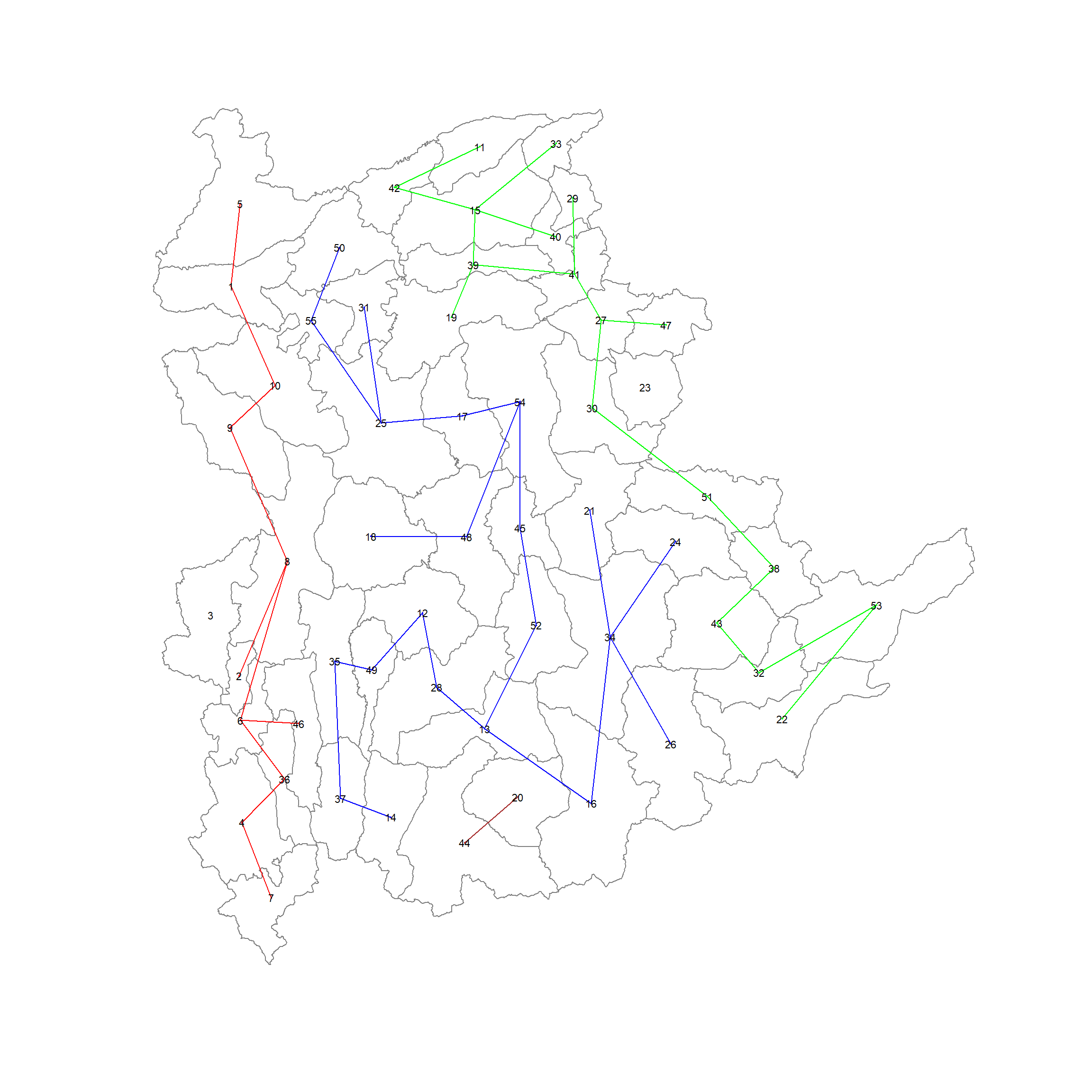
Visualising clusters in chloropeth map
groups_mat<- as.matrix(skater.clust6$groups)
shan_sf_spatialcluster <- cbind(shan_sf_cluster, as.factor(groups_mat)) %>%
rename(`skater_CLUSTER` = `as.factor.groups_mat.`)
qtm(shan_sf_spatialcluster, "skater_CLUSTER")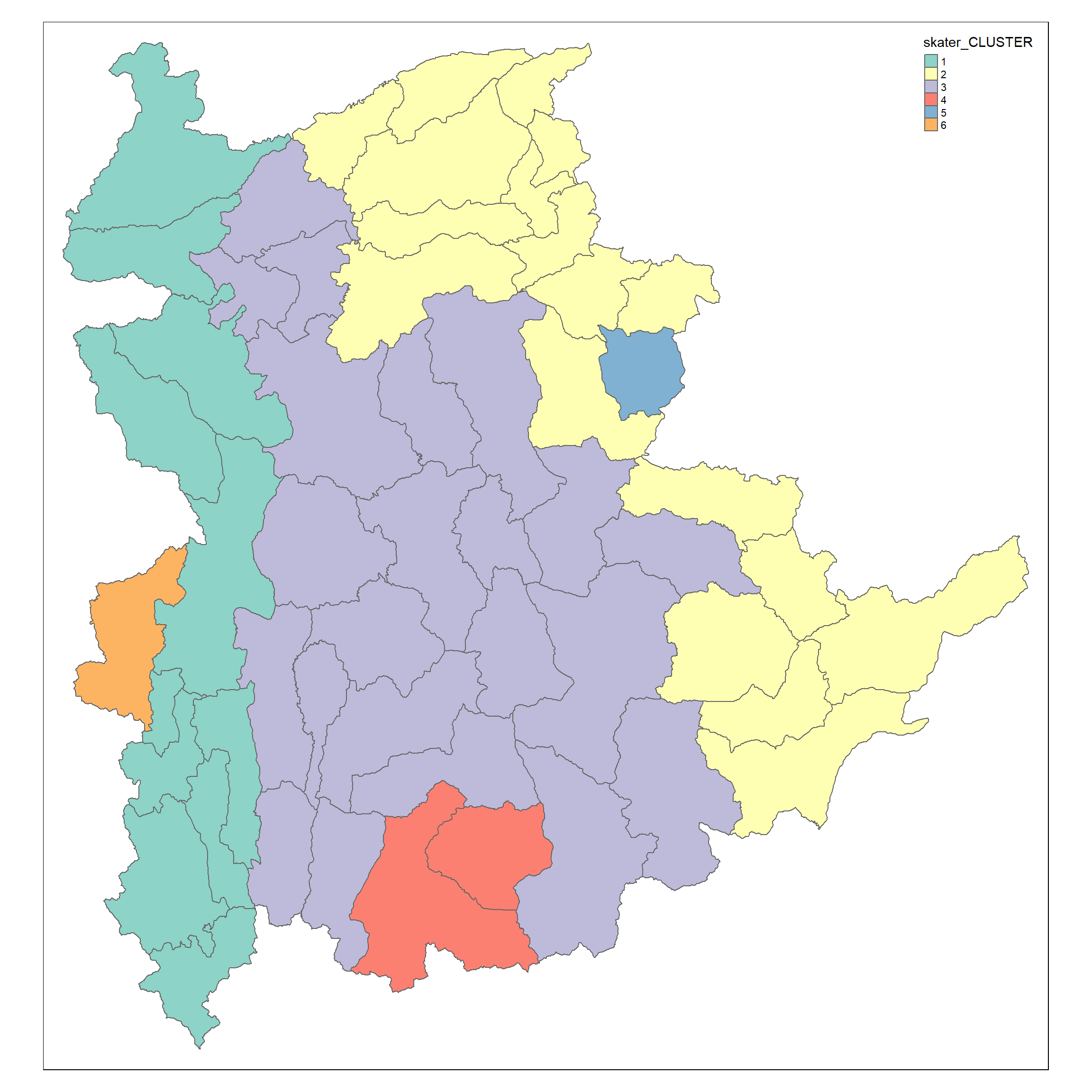
ClustGeo Algoritm
- Compute Spatial Distance Matrix To compute the distance matrix using st_distance() of sf package.
dist <- st_distance(shan_sf, shan_sf)
distmat <- as.dist(dist)- Cluster Graph
cr <- choicealpha(proxmat, distmat,
range.alpha = seq(0, 1, 0.1),
K=6, graph = TRUE)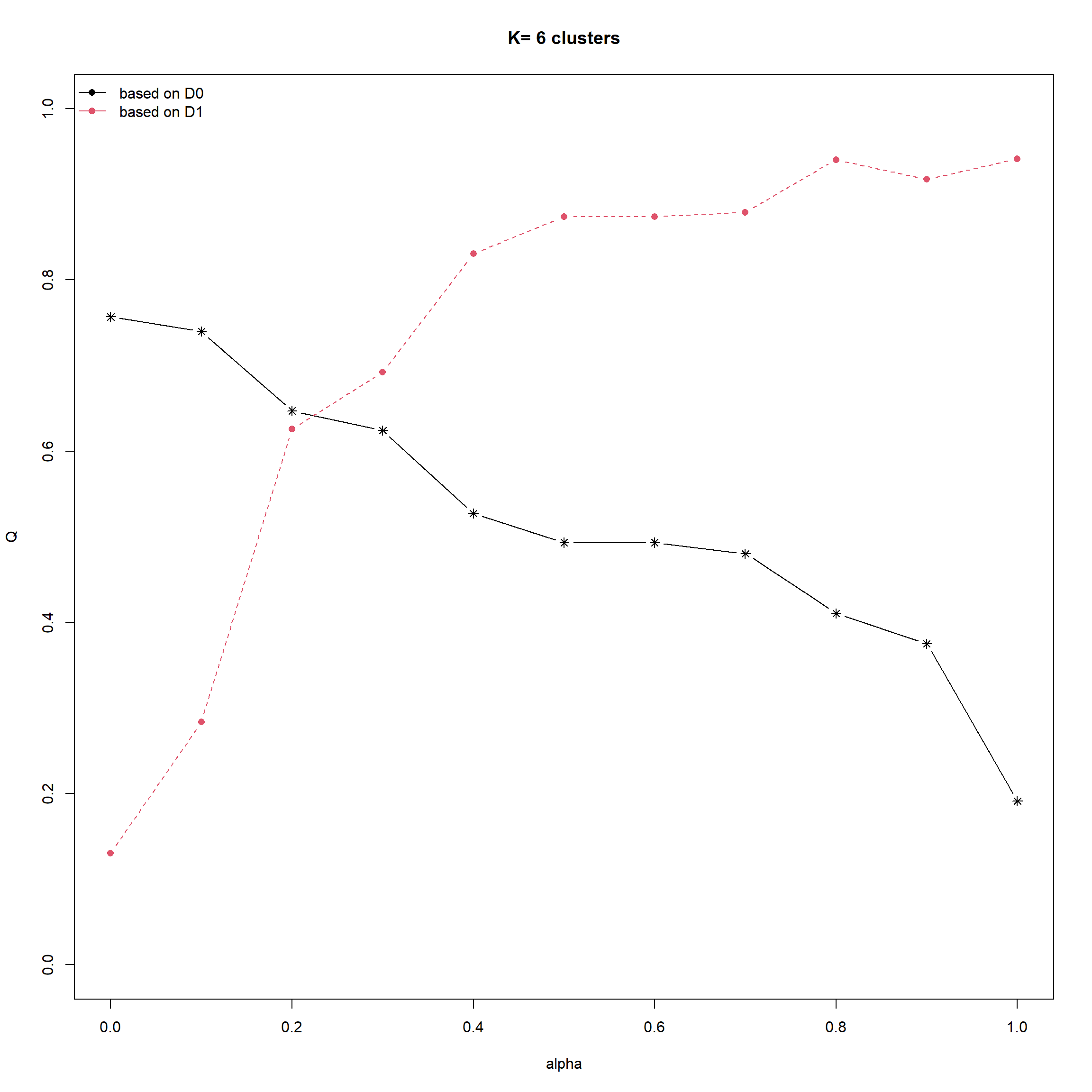
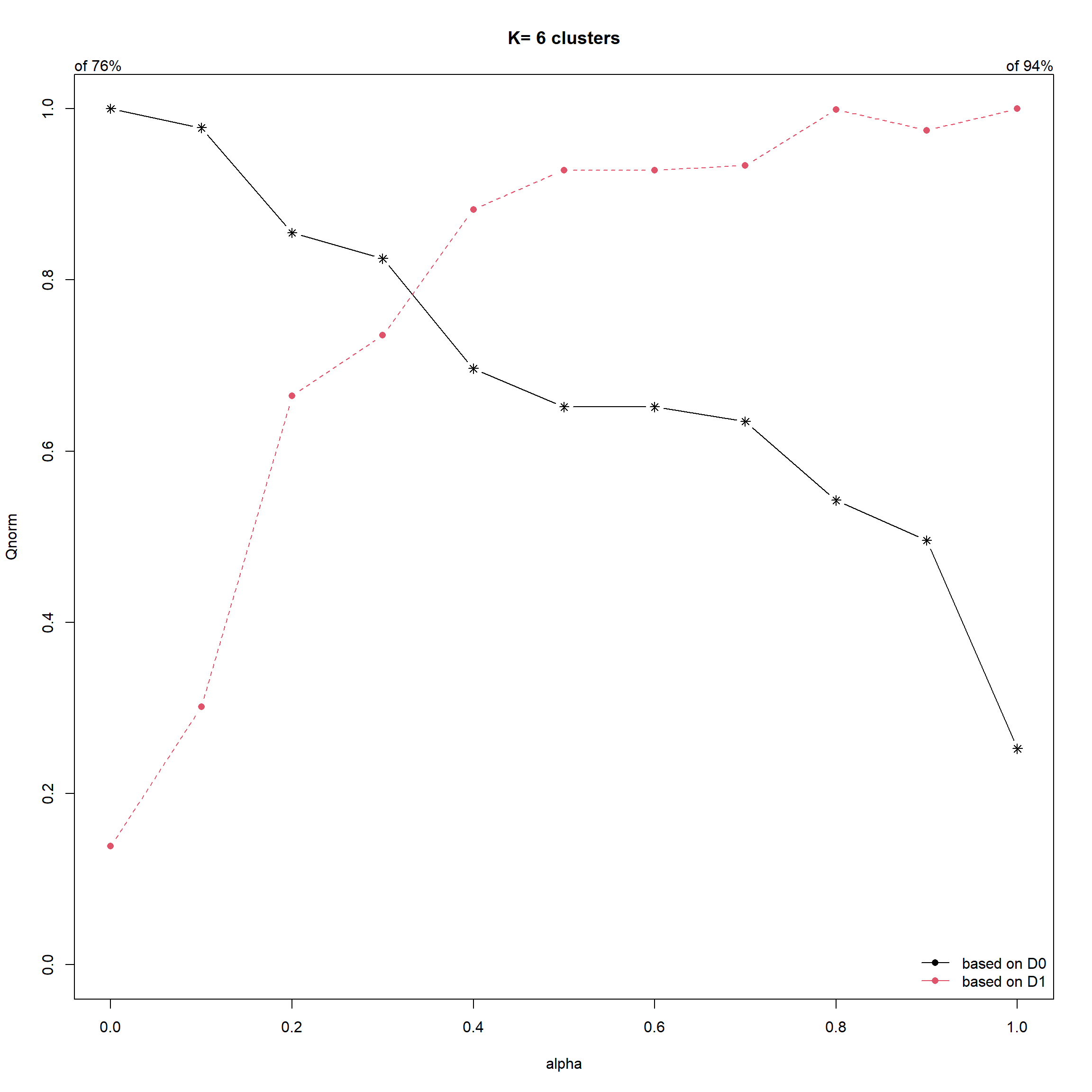
- Saving ClustGeo Output
clustG <- hclustgeo(proxmat, distmat, alpha = 0.2)
groups <- as.factor(cutree(clustG, k=6))
shan_sf_GclusterGeo <- cbind(shan_sf, as.matrix(groups)) %>%
rename(`clustGeo` = `as.matrix.groups.`)
qtm(shan_sf_GclusterGeo, "clustGeo")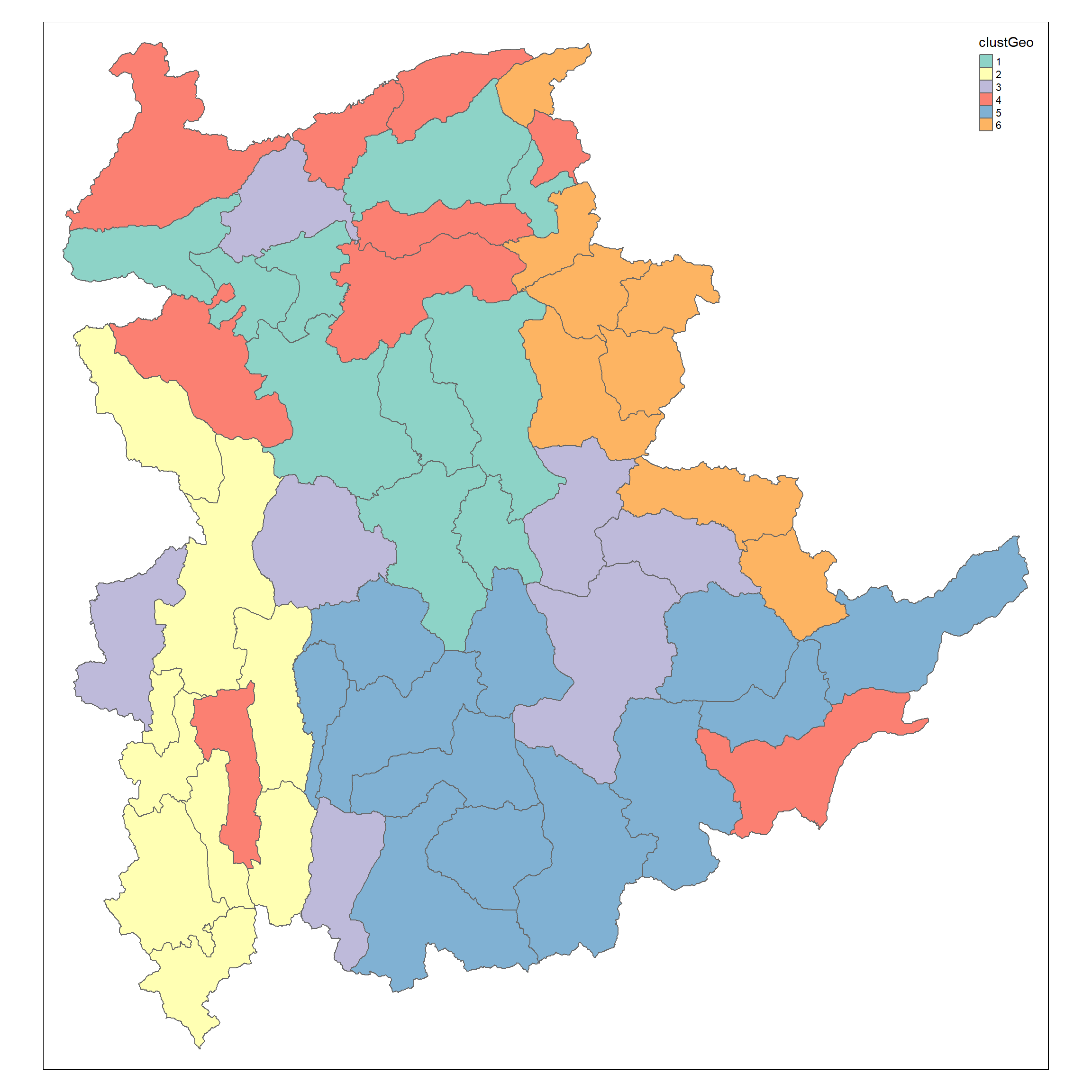
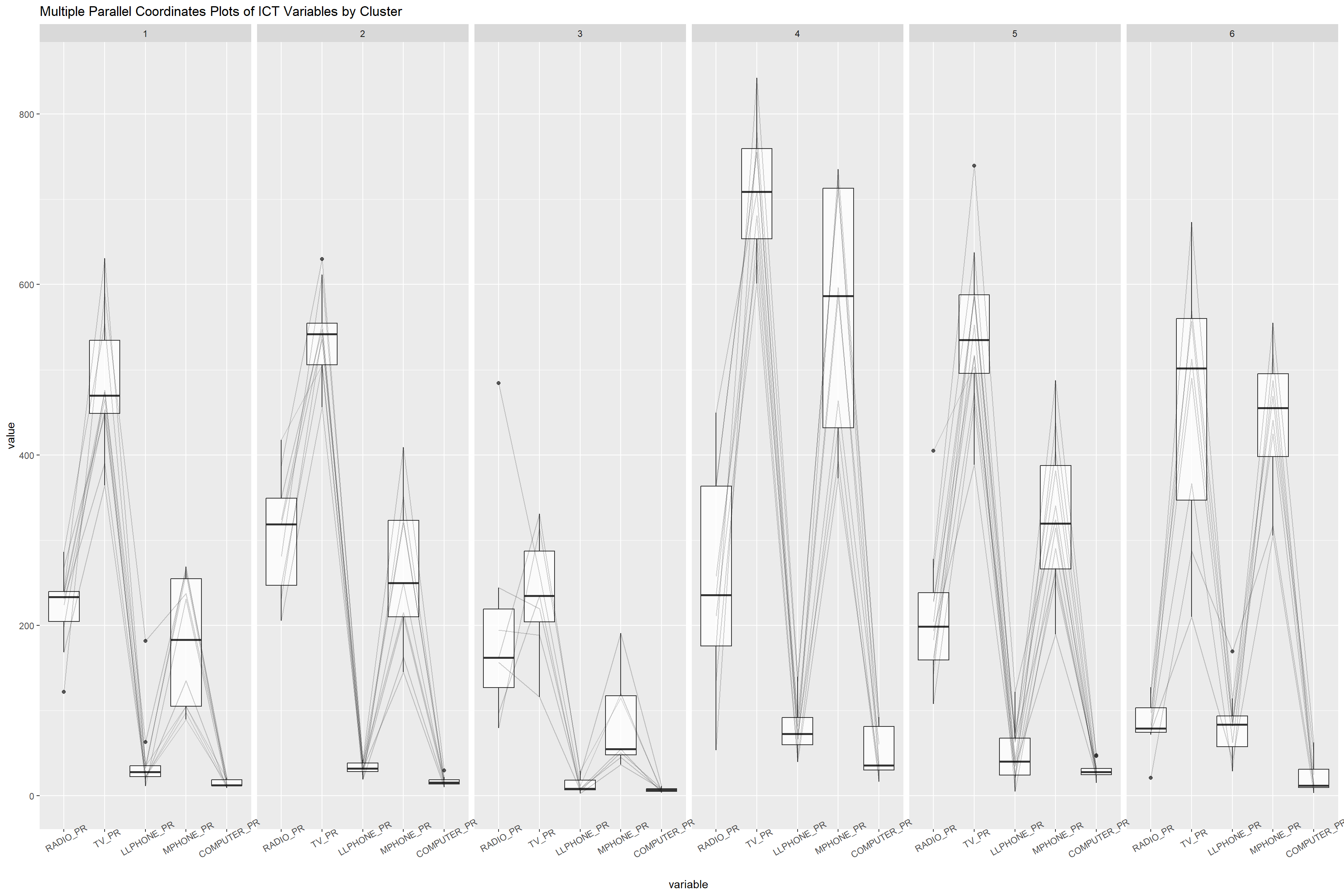
- Characterising the Clusters
ggparcoord(data = shan_sf_GclusterGeo,
columns = c(17:21),
scale = "globalminmax",
alphaLines = 0.2,
boxplot = TRUE,
title = "Multiple Parallel Coordinates Plots of ICT Variables by Cluster") +
facet_grid(~ clustGeo) +
theme(axis.text.x = element_text(angle = 30))